This web page was produced as an assignment for Genetics677, an undergraduate course at UW-Madison.
Protein motif/ domain
Motif
Following data retrieved from Motif
Following data retrieved from Motif
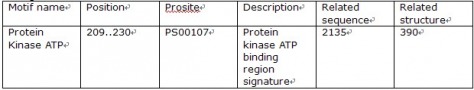
Using motif, I found only one motif from the protein of BMPR2. The motif name was “Protein kinase ATP.” As the name indicates, the region is responsible for APT to bind and to proceed its work.
Domain
Following data and figure retrieved from Pfam
Domain
Following data and figure retrieved from Pfam
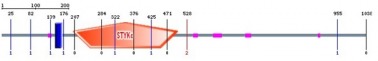
The green domain represents Activin receptor, while the long red block represents Protein kinase, which is an important domain for phosphorylation. The position of activin receptor is 58 to 131 and 203 to 501 for protein kinase. In addition, I received e-value and predicted active sites for both proteins. For activin receptor, the e-value is 47.0, and the predicted active site is 2.3e-12. I am not sure why the e-value is very huge like that number. I need to study further what the predicted active site means, which seems similar to e-value in this case. For protein kinase, the e-value is 158.1, and the predicted active site is 1.8e-16. Again, I need to understand what those numbers represent.
Following data and figure retrieved smart
Following data and figure retrieved smart
Name Begin End E-value
low complexity 132 142 -
Transmembrane 152 174 -
STYKc 203 502 9.71e-22
low complexity 545 558 -
low complexity 603 628 -
low complexity 694 710 -
low complexity 901 908 -
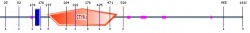
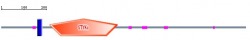
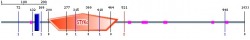
I liked the way of representing all data summary in Smart. As what I go from Pfam, the position, which is from 203 to 502. There are little variant of the end position (Pfam: 501 and Smart:502), so I am not sure which one is more accurate data. According to the data description, the blue block is tramsmembrane, and the pink is for low complexities. The orange pentagon with STKc is a protein kinase as what I saw from Pfam. The e-value for STKc is 9.71e-22. The e-value is only represented for STKc.
low complexity 132 142 -
Transmembrane 152 174 -
STYKc 203 502 9.71e-22
low complexity 545 558 -
low complexity 603 628 -
low complexity 694 710 -
low complexity 901 908 -
I liked the way of representing all data summary in Smart. As what I go from Pfam, the position, which is from 203 to 502. There are little variant of the end position (Pfam: 501 and Smart:502), so I am not sure which one is more accurate data. According to the data description, the blue block is tramsmembrane, and the pink is for low complexities. The orange pentagon with STKc is a protein kinase as what I saw from Pfam. The e-value for STKc is 9.71e-22. The e-value is only represented for STKc.
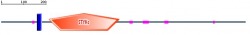
In addtion to look at human protein domain, I decided to search for domains in other species. As above figures are shown, BMPR2 protein domains are well conserved throughout species. Based on the length and location of each domain, Mus musculus (mouse) is a suitable animal model for my project to study further about BMPR2 signaling and how it is related to pulmonary arterial hypertension.
Following data and figure retrieved prosite
LKLLELIGRGRYGAVYKGSL--DERPVAVKVFSF-----ANRQNFINEKNIYRVpLmEHD
NIARFIVGDERVtadgrMEYLLVMEYYPNGSLCKYLSLH--TSDWVSSCRLAHSVTRGLA
YLHTElprgdhykpAISHRDLNSRNVLVKNDGTCVISDFGLSMRLtgnrlvRPGEEdnAA
ISEVGTIRYMAPEVLEgavnLRDCESAlkQVDMYALGLIYWEIFMrctdlfpGesvpeyQ
MAFQTEvgnhptfEDMQVLVSREKQR---PKFP--EAwkenslAVRSLKETIEDCWDQDA
EARLTAQCAEErMAEL
I also used prosite to find domain of my protein. As I expected the protein kinase domain that is for ATP biding site was found. The position of this domain is 203 to 504. I realized that the end position is slightly different from each website. The data also indicates the sequence of the region. Prosite gives scores rather than e-vlale. Scores for this domain is 35.623.
NIARFIVGDERVtadgrMEYLLVMEYYPNGSLCKYLSLH--TSDWVSSCRLAHSVTRGLA
YLHTElprgdhykpAISHRDLNSRNVLVKNDGTCVISDFGLSMRLtgnrlvRPGEEdnAA
ISEVGTIRYMAPEVLEgavnLRDCESAlkQVDMYALGLIYWEIFMrctdlfpGesvpeyQ
MAFQTEvgnhptfEDMQVLVSREKQR---PKFP--EAwkenslAVRSLKETIEDCWDQDA
EARLTAQCAEErMAEL
I also used prosite to find domain of my protein. As I expected the protein kinase domain that is for ATP biding site was found. The position of this domain is 203 to 504. I realized that the end position is slightly different from each website. The data also indicates the sequence of the region. Prosite gives scores rather than e-vlale. Scores for this domain is 35.623.